CSDB usage: symbol nomenclature for glycans (SNFG)
Glycan and glycoconjugate structures can be displayed using the recently agreed third version of the graphical notation published in Essentials of Glycobiology and called SNFG. The previous version of this notataion was also known as "CFG format". Below is a legend for monomeric residue symbols. More details are available at NCBI/Glycans/SNFG web page and in the dedicated publication: Varki et al. 2015, Glycobiology, 25(12):1323-1324 (DOI: 10.1093/glycob/cwv091).
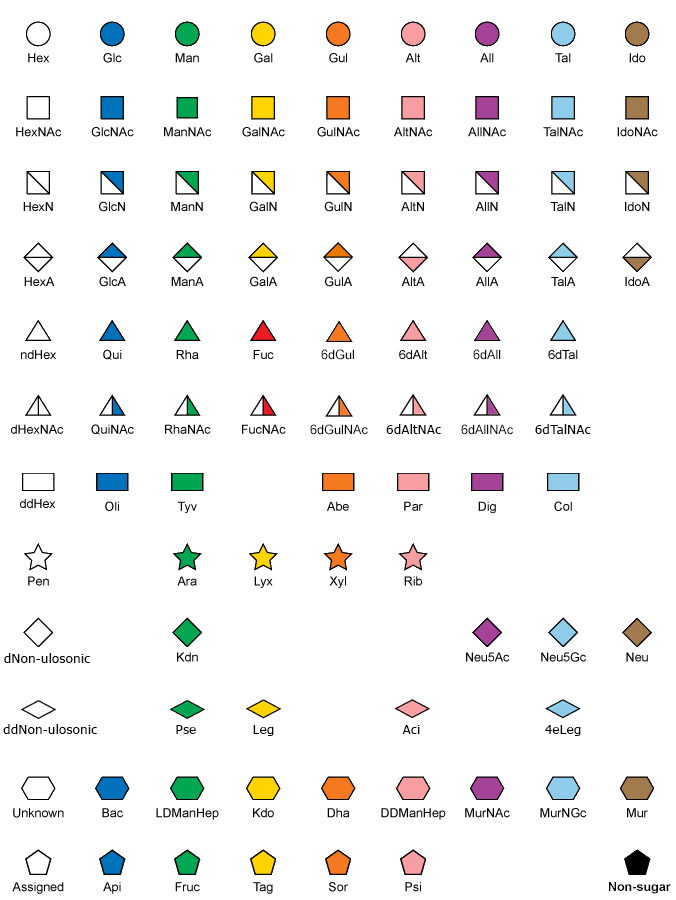
Passing the pointer over a symbol shows its full name. Clicking on a symbol links to the corresponding entry in MonosaccharideDB.
The following notes were adapted from the above publication, with some CSDB-specific features added:
- Each symbol represents a specific monosaccharide found in nature. Assigned symbols are shown in the Appendix to the Third Edition of Essentials of Glycobiology. To ensure harmony with prior publications, no changes were made to the Second Edition symbol set.
- Shapes and colors are completely consistent with stereochemistry only for hexoses, hexosamines, N-acetylhexosamines, hexuronates, and pentoses. Shapes only are consistent for deoxyhexoses, deoxy-N-acetylhexosamines, dideoxyhexoses, and nonulosonates.
- A symbol encodes a defined monosaccharide (including D or L) independently on rotation or mirroring. The divided diamonds for IdoA and AltA are inverted to indicate the commoner L-forms.
- White symbols based on the standard shapes designate monosaccharides with unknown stereochemistry (e.g., a white circle designates a hexose, type unknown). Colored pentagons and hexagons are used for other monosaccharides that are common in nature.
- Monosaccharides missing from this table are shown with a white shape derived from stereochemistry and a number inside; the legend of numbers is located below the figure.
- A white flat hexagon can be used for any unknown monosaccharide residue, for example for a heptose other than L-gro-D-manHep or D-gro-D-manHep.
- The commoner D-configuration and pyranose forms are assumed for all monosaccharides that do not have their absolute configuration implicit in their trivial name (Abe, Bac, Col, Dha, Kdn, Kdo, Mur, Neu, Par, Tyv), except for Ara, Fuc, Ido, Rha, Alt, Sor, and their derivatives, which are assumed to be in their commoner L-configuration, and Api, which is commonly in L-furanose form.
- Any deviations from the standard definitions above are labeled by letters iside the symbol (D or L for the rarely occurring absolute configuration; f, p or a for the uncommon ring size, o for alditols. Epimers at C8 of nonulosonates are labeled by 8D or 8L inside the symbol.
- Any modifications by monovalent substituents not implied by the monosaccharide name (such as GlcNAc or Neu5Ac) are listed as text (e.g. 3Me 4Ac) beside the residue icons.
- Linkage positions and anomericities are shown beside the links between residues. Double links mean that residues are linked twice (e.g. Pyr=4,6Gal). Dashed links mean that a donor residue is non-stoichimetric or not always present. Black links represent ether (incl., glycosidic), (di)ester, or amidic bonds; white links represent C-C bonds.
- All monosaccharide glycosidic linkages are assumed to originate from C-1 except for 2-ketoses, which are assumed to be linked from C-2.
- An internal sulfo- or phosphodiester is shown as -P- between the symbols for the linked residues.
- Enumerated black pentagons depict residues other than monosaccharides; the legend of numbers is located below the figure.
- [...] stands for previous or next repeating unit in regular polymeric structures.
- Alternative branches are listed vertically in grey boxes replacing a residue. The alternation logic (OR or XOR) is shown in white at the right of the grey box.
Example structure in SNFG notation:
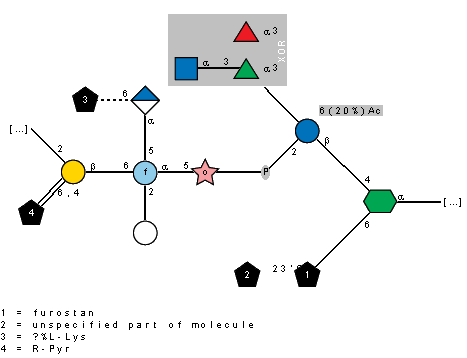
and in Sweet-DB notation for reference:
Subst1-(1C-23')-Subst-(1-6)-+
|
?%L-Lys-(2-6)-a-D-GlcpA-(1-5)-+ |
| |
R-Pyr-(2-6:2-4)-+ | /Variants 0/-+ |
| | | |
-2)-b-D-Galp-(1-6)-a-D-Talf-(1-5)-D-Rib-ol-(1--P--2)--b-D-Glcp6(20%)Ac-(1-4)-L-gro-a-D-manHepp-(1-
|
HEX-(1-2)-+
/Variants 0/ is:
a-L-Fucp-(1-3)-
OR (exclusively)
a-D-GlcpNAc-(1-3)-a-L-Rhap-(1-3)-
Subst = furostan;
Subst1 = unspecified part of molecule |