Taxonomic group: bacteria / Proteobacteria
(Phylum: Proteobacteria)
Associated disease: infection due to Escherichia coli [ICD11:
XN6P4 
]
The structure was elucidated in this paperNCBI PubMed ID: 7504582Publication DOI: 10.1016/0008-6215(93)84134-rJournal NLM ID: 0043535Publisher: Elsevier
Institutions: Institute for Biological Sciences, National Research Council of Canada, Ottawa, Ontario
The structure of the O-polysaccharide of the smooth lipopolysaccharide (LPS) produced by Escherichia coli O64:K99 was investigated by SDS-PAGE, composition, periodate oxidation, methylation, partial hydrolysis, and 1D and 2D nuclear magnetic resonance analyses, made on the native o-chain and its reduction and periodate degradation products. The E. coli O64 antigenic O-chain was found to be a high molecular weight glycan composed of D-galactose, D-glucuronic acid, 2-acetamido-2-deoxy-D-glucose, and 2-acetamido-2-deoxy-D-mannose (2:1:1:1) and was a polymer of branched pentasaccharide repeating units having the structure: [formula: see text]
Structure type: polymer chemical repeating unit
Location inside paper: abstract
Compound class: O-polysaccharide, O-antigen
Contained glycoepitopes: IEDB_115136,IEDB_135813,IEDB_136044,IEDB_137340,IEDB_137472,IEDB_1391962,IEDB_140630,IEDB_141794,IEDB_141807,IEDB_142078,IEDB_143794,IEDB_149549,IEDB_150899,IEDB_151531,IEDB_153201,IEDB_156493,IEDB_190606,IEDB_423153,SB_137,SB_165,SB_166,SB_187,SB_195,SB_29,SB_7,SB_88
Methods: 13C NMR, 1H NMR, NMR-2D
Related record ID(s): 108699
NCBI Taxonomy refs (TaxIDs): 562Reference(s) to other database(s): GTC:G07045HY, GlycomeDB:
28112
Show glycosyltransferases
NMR conditions: in D2O at 343 K
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6
3,3,3,2 Ac 173.4-175.2 22.7-23.2
3,3,3 aDManpN 100.2 50.9 76.4 72.4 73.1 61.1
3,3 bDGlcpA 104.5 72.5 81.8 65.5 75.4 ?
3,6 bDGalp 104.0 71.7 73.4 69.5 76.0 61.8
3 bDGalp 104.2 70.5 83.0 69.0 76.0 70.0
2 Ac 173.4-175.2 22.7-23.2
bDGlcpN 99.5 55.0 84.4 69.6 76.0 61.4
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6
3,3,3,2 Ac - 1.988-2.035
3,3,3 aDManpN 5.231 4.489 4.226 3.782 4.056 3.841-3.841
3,3 bDGlcpA 4.760 3.529 3.697 3.801 3.990 -
3,6 bDGalp 4.449 3.548 3.648 3.936 3.684 3.767-3.789
3 bDGalp 4.477 3.706 3.799 4.156 3.919 3.909-4.053
2 Ac - 1.988-2.035
bDGlcpN 4.665 3.857 3.804 3.579 3.515 3.773-3.931
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6
3,3,3,2 Ac 22.7-23.2/1.988-2.035
3,3,3 aDManpN 100.2/5.231 50.9/4.489 76.4/4.226 72.4/3.782 73.1/4.056 61.1/3.841-3.841
3,3 bDGlcpA 104.5/4.760 72.5/3.529 81.8/3.697 65.5/3.801 75.4/3.990
3,6 bDGalp 104.0/4.449 71.7/3.548 73.4/3.648 69.5/3.936 76.0/3.684 61.8/3.767-3.789
3 bDGalp 104.2/4.477 70.5/3.706 83.0/3.799 69.0/4.156 76.0/3.919 70.0/3.909-4.053
2 Ac 22.7-23.2/1.988-2.035
bDGlcpN 99.5/4.665 55.0/3.857 84.4/3.804 69.6/3.579 76.0/3.515 61.4/3.773-3.931
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 |
|---|
| 3,3,3,2 | Ac |
| 1.988
2.035 | |
| 3,3,3 | aDManpN | 5.231 | 4.489 | 4.226 | 3.782 | 4.056 | 3.841
3.841 |
| 3,3 | bDGlcpA | 4.760 | 3.529 | 3.697 | 3.801 | 3.990 |
|
| 3,6 | bDGalp | 4.449 | 3.548 | 3.648 | 3.936 | 3.684 | 3.767
3.789 |
| 3 | bDGalp | 4.477 | 3.706 | 3.799 | 4.156 | 3.919 | 3.909
4.053 |
| 2 | Ac |
| 1.988
2.035 | |
| | bDGlcpN | 4.665 | 3.857 | 3.804 | 3.579 | 3.515 | 3.773
3.931 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 |
|---|
| 3,3,3,2 | Ac | 173.4
175.2 | 22.7
23.2 | |
| 3,3,3 | aDManpN | 100.2 | 50.9 | 76.4 | 72.4 | 73.1 | 61.1 |
| 3,3 | bDGlcpA | 104.5 | 72.5 | 81.8 | 65.5 | 75.4 | ? |
| 3,6 | bDGalp | 104.0 | 71.7 | 73.4 | 69.5 | 76.0 | 61.8 |
| 3 | bDGalp | 104.2 | 70.5 | 83.0 | 69.0 | 76.0 | 70.0 |
| 2 | Ac | 173.4
175.2 | 22.7
23.2 | |
| | bDGlcpN | 99.5 | 55.0 | 84.4 | 69.6 | 76.0 | 61.4 |
|
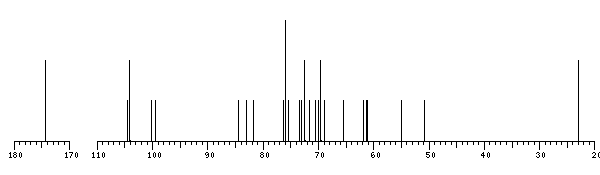
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error Found 1 record.
Displayed record 1
Found 1 record.
Displayed record 1
 report error
report error
 ]
]