Taxonomic group: bacteria / Proteobacteria
(Phylum: Proteobacteria)
Associated disease: infection due to Acinetobacter baumannii [ICD11:
XN8LS 
]
The structure was elucidated in this paperNCBI PubMed ID: 25917131Publication DOI: 10.1016/j.carres.2015.03.016Journal NLM ID: 0043535Publisher: Elsevier
Correspondence: C. De Castro <decastro

unina.it>
Institutions: School of Molecular Bioscience, The University of Sydney, Sydney, NSW 2006, Australia, Department of Chemical Sciences, Complesso Universitario Monte Sant'Angelo, Napoli, Italy
The structure of the capsular polysaccharide (CPS) from an Acinetobacter baumannii global clone 2 (GC2) clinical isolate RBH4 that carries the KL6 gene cluster was elucidated by means of chemical and spectroscopical methods. The repeating unit of K6 CPS is linear and contains N-acetyl-D-galactosamine (D-GalpNAc), two D-galactose (D-Galp) residues and 5,7-di-N-acetylpseudaminic acid (Pse5Ac7Ac). The synthesis of these sugars could be attributed to genes in the KL6 capsule biosynthesis gene cluster, and the formation of the linkages between the sugars were assigned to glycosyltransferases or the Wzy polymerase encoded in KL6.
Acinetobacter baumannii, capsular polysaccharide, NMR spectroscopy, pseudaminic acid, K locus, KL6 gene cluster
Structure type: polymer chemical repeating unit
Location inside paper: p.33, fig.5, p.34, fig.6, K6
Compound class: CPS
Contained glycoepitopes: IEDB_130648,IEDB_134627,IEDB_136044,IEDB_137472,IEDB_137473,IEDB_1391961,IEDB_1391963,IEDB_141584,IEDB_141794,IEDB_143260,IEDB_153201,IEDB_156493,IEDB_190606,IEDB_885822,SB_165,SB_166,SB_187,SB_195,SB_23,SB_24,SB_7,SB_8,SB_88
Methods: 13C NMR, 1H NMR, methylation, NMR-2D, GC-MS, sugar analysis, enzymatic hydrolysis, extraction, acetylation, bioinformatic analysis, dialysis
Biosynthesis and genetic data: genetic data
Related record ID(s): 30766
NCBI Taxonomy refs (TaxIDs): 470Reference(s) to other database(s): GTC:G18298UG
Show glycosyltransferases
NMR conditions: in D2O at 310 K
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9
3,6,6,5 Ac 175.3 ?
3,6,6,7 Ac 174.9 ?
3,6,6 bXPsep ? ? 34.4 73.1 48.8 74.8 54.9 70.2 18.3
3,6 bDGalp 105.8 72.0 73.9 70.1 74.8 65.2
3 bDGalp 105.1 72.0 73.9 70.1 75.1 71.0
2 Ac
aDGalpN 96.7 49.9 78.9 69.9 72.3 62.9
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6 H7 H8 H9
3,6,6,5 Ac
3,6,6,7 Ac
3,6,6 bXPsep - - 1.63-2.56 3.86 4.29 4.06 4.01 4.10 1.21
3,6 bDGalp 4.42 3.53 3.64 3.95 3.79 3.60-3.93
3 bDGalp 4.47 3.52 3.64 3.96 3.84 3.81-4.11
2 Ac
aDGalpN 5.04 4.27 3.85 4.24 4.18 3.76-3.76
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6 C7/H7 C8/H8 C9/H9
3,6,6,5 Ac NMR TSV error 2: unequal length of 13C and 1H datasets
3,6,6,7 Ac NMR TSV error 2: unequal length of 13C and 1H datasets
3,6,6 bXPsep 34.4/1.63-2.56 73.1/3.86 48.8/4.29 74.8/4.06 54.9/4.01 70.2/4.10 18.3/1.21
3,6 bDGalp 105.8/4.42 72.0/3.53 73.9/3.64 70.1/3.95 74.8/3.79 65.2/3.60-3.93
3 bDGalp 105.1/4.47 72.0/3.52 73.9/3.64 70.1/3.96 75.1/3.84 71.0/3.81-4.11
2 Ac
aDGalpN 96.7/5.04 49.9/4.27 78.9/3.85 69.9/4.24 72.3/4.18 62.9/3.76-3.76
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 | H7 | H8 | H9 |
|---|
| 3,6,6,5 | Ac | |
| 3,6,6,7 | Ac | |
| 3,6,6 | bXPsep |
|
| 1.63
2.56 | 3.86 | 4.29 | 4.06 | 4.01 | 4.10 | 1.21 |
| 3,6 | bDGalp | 4.42 | 3.53 | 3.64 | 3.95 | 3.79 | 3.60
3.93 | |
| 3 | bDGalp | 4.47 | 3.52 | 3.64 | 3.96 | 3.84 | 3.81
4.11 | |
| 2 | Ac | |
| | aDGalpN | 5.04 | 4.27 | 3.85 | 4.24 | 4.18 | 3.76
3.76 | |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 |
|---|
| 3,6,6,5 | Ac | 175.3 | ? | |
| 3,6,6,7 | Ac | 174.9 | ? | |
| 3,6,6 | bXPsep | ? | ? | 34.4 | 73.1 | 48.8 | 74.8 | 54.9 | 70.2 | 18.3 |
| 3,6 | bDGalp | 105.8 | 72.0 | 73.9 | 70.1 | 74.8 | 65.2 | |
| 3 | bDGalp | 105.1 | 72.0 | 73.9 | 70.1 | 75.1 | 71.0 | |
| 2 | Ac | |
| | aDGalpN | 96.7 | 49.9 | 78.9 | 69.9 | 72.3 | 62.9 | |
|
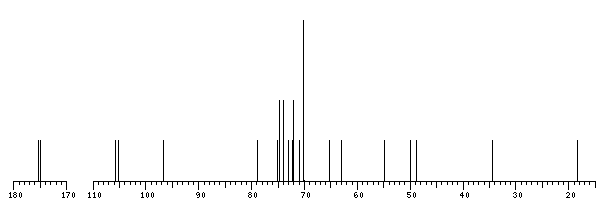
The spectrum also has 4 signals at unknown positions (not plotted). |
There is only one chemically distinct structure:
 report error
report error Found 1 record.
Displayed record 1
Found 1 record.
Displayed record 1
 report error
report error
 ]
] unina.it>
unina.it>