Taxonomic group: bacteria / Proteobacteria
(Phylum: Proteobacteria)
Host organism: Homo sapiens
Associated disease: infection due to Acinetobacter baumannii [ICD11:
XN8LS 
]
The structure was elucidated in this paperNCBI PubMed ID: 26432612Publication DOI: 10.1016/j.carres.2015.09.004Journal NLM ID: 0043535Publisher: Elsevier
Correspondence: Y.A. Knirel <yknirel

gmail.com>
Institutions: N.D. Zelinsky Institute of Organic Chemistry, Russian Academy of Sciences, Moscow, Russia, M. M. Shemyakin & Y. A. Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, Moscow, Russia, State Research Center for Applied Microbiology and Biotechnology, Obolensk, Moscow Region, Russia, School of Molecular Bioscience, The University of Sydney, Sydney, Australia, School of Biomedical Sciences, Queensland University of Technology, Brisbane, Australia, Moscow Institute of Physics and Technology, Dolgoprudny, Moscow Region, Russia
Neutral capsular polysaccharides (CPSs) were isolated from Acinetobacter baumannii NIPH190, NIPH201, and NIPH615. The CPSs were found to contain common monosaccharides only and to be branched with a side-chain 1→3-linked β-d-glucopyranose residue. Structures of the oligosaccharide repeat units (K units) of the CPSs were elucidated by 1D and 2D 1H and 13C NMR spectroscopy. Novel CPS biosynthesis gene clusters, designated KL30, KL45, and KL48, were found at the K locus in the genome sequences of NIPH190, NIPH201, and NIPH615, respectively. The genetic content of each gene cluster correlated with the structure of the CPS unit established, and therefore, the capsular types of the strains studied were designated as K30, K45, and K48, respectively. The initiating sugar of each K unit was predicted, and glycosyltransferases encoded by each gene cluster were assigned to the formation of the linkages between sugars in the corresponding K unit.
Acinetobacter baumannii, capsular polysaccharide structure, glycosyltransferase, K locus, polysaccharide gene cluster
Structure type: polymer chemical repeating unit
Location inside paper: p.82, table 2, p.84, chart 1, K45 (NIPH201)
Compound class: CPS
Contained glycoepitopes: IEDB_130648,IEDB_137340,IEDB_137473,IEDB_1391961,IEDB_141584,IEDB_141807,IEDB_142488,IEDB_146664,IEDB_151531,IEDB_885822,IEDB_983931,SB_192
Methods: 13C NMR, 1H NMR, NMR-2D, sugar analysis, GLC, de-O-acetylation, NMR-1D, GPC, extraction, bioinformatic analysis
Biosynthesis and genetic data: genetic data
Comments, role: CPS from A. baumannii K45 (NIPH201).
Related record ID(s): 30676, 30877, 30878
NCBI Taxonomy refs (TaxIDs): 1217630Reference(s) to other database(s): GTC:G57327XB
Show glycosyltransferases
NMR conditions: in D2O at 293 K
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6
3,4,2 Ac 175.5-175.8 23.4-23.7
3,4,6 50%Ac ? 21.4
3,4 aDGalpN 98.3 51. 67.7 77.6 69.0 61.6
3,2 Ac 175.4-175.6 23.0-23.6
3,3 bDGlcp 106.3 74.3 76.8 71.0 77.0 62.0
3 aDGalpN 99.0 50.0 77.6 75.1 72.8 60.7
2 Ac 175.4-175.6 23.0-23.6
aDGlcpN 99.9 53.7 77.0 71.7 73.2 60.8
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6
3,4,2 Ac - 2.04-2.06
3,4,6 50%Ac - 2.12
3,4 aDGalpN 5.08 4.26 4.18 4.12 4.72 4.07-4.19
3,2 Ac - 2.04-2.06
3,3 bDGlcp 4.45 3.09 3.47 3.21 3.38 3.70-3.89
3 aDGalpN 5.42 4.49 3.89 4.32 3.96 3.65-3.71
2 Ac - 2.04-2.06
aDGlcpN 4.79 4.10 3.99 3.79 4.16 3.72-3.80
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6
3,4,2 Ac 23.4-23.7/2.04-2.06
3,4,6 50%Ac 21.4/2.12
3,4 aDGalpN 98.3/5.08 51./4.26 67.7/4.18 77.6/4.12 69.0/4.72 61.6/4.07-4.19
3,2 Ac 23.0-23.6/2.04-2.06
3,3 bDGlcp 106.3/4.45 74.3/3.09 76.8/3.47 71.0/3.21 77.0/3.38 62.0/3.70-3.89
3 aDGalpN 99.0/5.42 50.0/4.49 77.6/3.89 75.1/4.32 72.8/3.96 60.7/3.65-3.71
2 Ac 23.0-23.6/2.04-2.06
aDGlcpN 99.9/4.79 53.7/4.10 77.0/3.99 71.7/3.79 73.2/4.16 60.8/3.72-3.80
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 |
|---|
| 3,4,2 | Ac |
| 2.04
2.06 | |
| 3,4,6 | 50%Ac |
| 2.12 | |
| 3,4 | aDGalpN | 5.08 | 4.26 | 4.18 | 4.12 | 4.72 | 4.07
4.19 |
| 3,2 | Ac |
| 2.04
2.06 | |
| 3,3 | bDGlcp | 4.45 | 3.09 | 3.47 | 3.21 | 3.38 | 3.70
3.89 |
| 3 | aDGalpN | 5.42 | 4.49 | 3.89 | 4.32 | 3.96 | 3.65
3.71 |
| 2 | Ac |
| 2.04
2.06 | |
| | aDGlcpN | 4.79 | 4.10 | 3.99 | 3.79 | 4.16 | 3.72
3.80 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 |
|---|
| 3,4,2 | Ac | 175.5
175.8 | 23.4
23.7 | |
| 3,4,6 | 50%Ac | ? | 21.4 | |
| 3,4 | aDGalpN | 98.3 | 51. | 67.7 | 77.6 | 69.0 | 61.6 |
| 3,2 | Ac | 175.4
175.6 | 23.0
23.6 | |
| 3,3 | bDGlcp | 106.3 | 74.3 | 76.8 | 71.0 | 77.0 | 62.0 |
| 3 | aDGalpN | 99.0 | 50.0 | 77.6 | 75.1 | 72.8 | 60.7 |
| 2 | Ac | 175.4
175.6 | 23.0
23.6 | |
| | aDGlcpN | 99.9 | 53.7 | 77.0 | 71.7 | 73.2 | 60.8 |
|
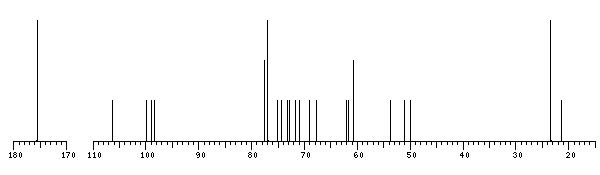
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error Found 1 record.
Displayed record 1
Found 1 record.
Displayed record 1
 report error
report error
 ]
] gmail.com>
gmail.com>