Taxonomic group: fungi / Ascomycota
(Phylum: Ascomycota)
The structure was elucidated in this paperNCBI PubMed ID: 15470229Publication DOI: 10.1093/glycob/cwi002Journal NLM ID: 9104124Publisher: IRL Press at Oxford University Press
Correspondence: f.hochstenbach

amc.uva.nl
Institutions: Bijvoet Center, Department of Bio-Organic Chemistry, Section of Glycoscience and Biocatalysis, Utrecht University, Utrecht, the Netherlands, Swammerdam Institute for Life Sciences, University of Amsterdam, Nieuwe Achtergracht 166, 1018 WV Amsterdam, The Netherlands, Department of Medical Biochemistry, Academic Medical Center, University of Amsterdam, Meibergdreef 15, 1105 AZ Amsterdam, The Netherlands, Department of Molecular Cell Biology, Institute for Biomembranes, Utrecht University, Utrecht, The Netherlands, Laboratory for Molecular Biology of Plants, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Kerklaan 30, 9715 NN Haren, The Netherlands
Morphology and structural integrity of fungal cells depend on cell wall polysaccharides. The chemical structure and biosynthesis of two types of these polysaccharides, chitin and (1→3)-β-glucan, have been studied extensively, whereas little is known about α-glucan. Here we describe the chemical structure of α-glucan isolated from wild-type and mutant cell walls of the fission yeast Schizosaccharomyces pombe. Wild-type α-glucan was found to consist of a single population of linear glucose polymers, approximately 260 residues in length. These glucose polymers were composed of two interconnected linear chains, each consisting of approximately 120 (1→3)-linked α-d-glucose residues and some (1→4)-linked α-D-glucose residues at the reducing end. By contrast, α-glucan of an α-glucan synthase mutant with an aberrant cell morphology and reduced α-glucan levels consisted of a single chain only. We propose that α-glucan biosynthesis involves an ordered series of events, whereby two α-glucan chains are coupled to create mature cell wall α-glucan. This mature form of cell wall α-glucan is essential for fission-yeast morphogenesis.
cell wall, polysaccharides, morphogenesis, Schizosaccharomyces pombe, alpha-glucan, fission yeast
Structure type: fragment of a bigger structure
Location inside paper: abstract, p.249, fig.3, part B
Trivial name: α-D-glucan, glycogen, amylose-like α-D-glucan
Compound class: EPS, cell wall polysaccharide
Contained glycoepitopes: IEDB_140629,IEDB_142488,IEDB_144998,IEDB_146664,IEDB_420417,IEDB_420418,IEDB_420421,IEDB_857742,IEDB_983931,SB_192
Methods: 13C NMR, 1H NMR, methylation, periodate oxidation, NMR-2D, GC-MS, acid hydrolysis, alkaline degradation, GC, Smith degradation, composition analysis, HPSEC, enzymatic digestion, acetylation, transmission electron microscopy
Comments, role: Part of cell wall, minor component B (see ID: 41279) . (1-4)-linked a-glucose residues are present at the reducing end of a-glucan.
Related record ID(s): 41279, 49051
NCBI Taxonomy refs (TaxIDs): 4896Reference(s) to other database(s): GTC:G05740LL
Show glycosyltransferases
NMR conditions: in DMSO-d6 at 353 K
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6
aDGlcp 99.8 71.7 72.9 78.5 71.2 ?
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6
aDGlcp 5.133 3.352 3.714 3.390 3.652 3.612-3.701
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6
aDGlcp 99.8/5.133 71.7/3.352 72.9/3.714 78.5/3.390 71.2/3.652 ?/3.612-3.701
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 |
|---|
| | aDGlcp | 5.133 | 3.352 | 3.714 | 3.390 | 3.652 | 3.612
3.701 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 |
|---|
| | aDGlcp | 99.8 | 71.7 | 72.9 | 78.5 | 71.2 | ? |
|
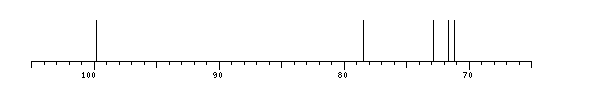
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error Found 1 record.
Displayed record 1
Found 1 record.
Displayed record 1
 report error
report error
 amc.uva.nl
amc.uva.nl