Taxonomic group: fungi / Mucoromycota
(Phylum: Mucoromycota)
NCBI PubMed ID: 30755333Publication DOI: 10.1016/j.bioorg.2019.01.046Journal NLM ID: 1303703Publisher: Orlando, FL: Academic Press
Correspondence: Sordon S <sandra.sordon

upwr.edu.pl>; Popłoński J <jaroslaw.poplonski

upwr.edu.pl>; Tronina T <tomasz.tronina

upwr.edu.pl>; Huszcza E <ewa.huszcza

upwr.edu.pl>
Institutions: Department of Chemistry, Wrocław University of Environmental and Life Sciences, Wrocław, Poland
In the present study, the species: Beauveria bassiana, Absidia coerulea and Absidia glauca were used in biotransformation of flavones (chrysin, apigenin, luteolin, diosmetin) and flavanones (pinocembrin, naringenin, eriodictyol, hesperetin). The Beauveria bassiana AM 278 strain catalyzed the methylglucose attachment reactions to the flavonoid molecule at positions C7 and C3'. The application of the Absidia genus (A. coerulea AM 93, A. glauca AM 177) as the biocatalyst resulted in the formation of glucosides with a sugar molecule present at C7 and C3' positions of flavonoids skeleton. Nine of obtained products have not been previously reported in the literature.
biotransformation, Beauveria bassiana, Absidia coerulea, microbial glycosylation, Absidia glauca, flavanones, flavones
Structure type: monomer
Location inside paper: Fig. 2, 12, supplementary materials, NMR data, 12
Trivial name: cosmosiin, apigenin 7-O-glucoside, rhoifolin, apigenin 7-glucoside, apigetrin
Compound class: glycoside, flavonoid glycoside, flavonol glycoside
Contained glycoepitopes: IEDB_142488,IEDB_146664,IEDB_613414,IEDB_983931,SB_192
Methods: 13C NMR, 1H NMR, NMR-2D, TLC, HPLC, biosynthetic methods, extraction, cell growth, HR-ESI-MS
Comments, role: NMR temperature was not specified
Related record ID(s): 49674, 49675, 49676, 49677, 49678, 49679, 49680, 49681, 49682, 49683, 49684, 49685, 49686, 49688, 49689, 49690, 49691, 49692
NCBI Taxonomy refs (TaxIDs): 90261,
4829Reference(s) to other database(s): CCSD:
50502, CBank-STR:1305
Show glycosyltransferases
NMR conditions: in DMSO-d6
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9 C10 C11 C12 C13 C14 C15 C16
7 bDGlcp 99.9 73.1 76.4 69.5 77.2 60.6
xXApigenin - 164.3 103.1 182.0 156.9 94.8 163.0 99.5 161.1 105.3 121.0 128.6 116.0 161.4 116.0 128.6
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6 H7 H8 H9 H10 H11 H12 H13 H14 H15 H16
7 bDGlcp 5.07 3.26 3.30 3.18 3.44 3.48-3.71
xXApigenin - - 6.87 - - 6.83 - 6.44 - - - 7.96 6.94 - 6.94 7.96
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6 C7/H7 C8/H8 C9/H9 C10/H10 C11/H11 C12/H12 C13/H13 C14/H14 C15/H15 C16/H16
7 bDGlcp 99.9/5.07 73.1/3.26 76.4/3.30 69.5/3.18 77.2/3.44 60.6/3.48-3.71
xXApigenin 103.1/6.87 94.8/6.83 99.5/6.44 128.6/7.96 116.0/6.94 116.0/6.94 128.6/7.96
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 | H7 | H8 | H9 | H10 | H11 | H12 | H13 | H14 | H15 | H16 |
|---|
| 7 | bDGlcp | 5.07 | 3.26 | 3.30 | 3.18 | 3.44 | 3.48
3.71 | |
| | xXApigenin |
|
| 6.87 |
|
| 6.83 |
| 6.44 |
|
|
| 7.96 | 6.94 |
| 6.94 | 7.96 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 | C10 | C11 | C12 | C13 | C14 | C15 | C16 |
|---|
| 7 | bDGlcp | 99.9 | 73.1 | 76.4 | 69.5 | 77.2 | 60.6 | |
| | xXApigenin |
| 164.3 | 103.1 | 182.0 | 156.9 | 94.8 | 163.0 | 99.5 | 161.1 | 105.3 | 121.0 | 128.6 | 116.0 | 161.4 | 116.0 | 128.6 |
|
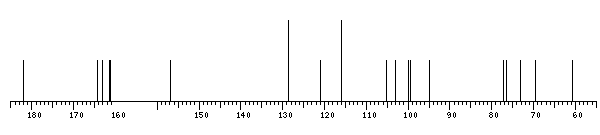
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error report error
report error
