Taxonomic group: bacteria / Actinobacteria
(Phylum: Actinobacteria)
Associated disease: infection due to Mycobacterium tuberculosis [ICD11:
XN1N2 
];
infection due to Mycobacterium bovis [ICD11:
XN8AB 
]
The structure was elucidated in this paperNCBI PubMed ID: 12063260Publication DOI: 10.1074/jbc.M204398200Journal NLM ID: 2985121RPublisher: Baltimore, MD: American Society for Biochemistry and Molecular Biology
Correspondence: laurent.kremer

ibl.fr
Institutions: Laboratoire de Glycobiologie Structurale et Fonctionnelle, CNRS UMR8576, Universite des Sciences et Technologies de Lille, F-59655 Villeneuve Ascq Cedex, France, School of Biosciences, University of Birmingham, Edgbaston, Birmingham, B15 2TT United Kingdom, Laboratoire des Mecanismes Moleculaires de la Pathogenie
Lipomannan (LM) and lipoarabinomannan (LAM) are major glycolipids present in the mycobacterial cell wall that are able to modulate the host immune response. In this study, we have undertaken the structural determination of these important modulins in Mycobacterium chelonae, a fast growing pathogenic mycobacterial species. One-dimensional and two-dimensional NMR spectra were used to demonstrate that LM and LAM from M. chelonae, designated CheLM and CheLAM, respectively, possess structures that differ from the ones reported earlier in other mycobacterial species. Analysis by gas chromatography/mass spectrometry of the phosphatidyl-myo-inositol anchor, which is thought to play a role in the biological functions of these lipoglycans, pointed to a high degree of heterogeneity based on numerous combinations of acyl groups on the C-1 and C-2 positions of the glycerol moiety. Characterization of the mannan core of CheLM and CheLAM revealed the presence of novel α1,3-mannopyranosyl side chains. This motif, which reacted specifically with the lectin from Galanthus nivalis, was found to be unique among a panel of nine mycobacterial species. Then, CheLM and CheLAM were found to be devoid of both the mannooligosaccharide cap present in Mycobacterium tuberculosis and the inositol phosphate cap present in Mycobacterium smegmatis and other fast growing species. Tumor necrosis factor-alpha and interleukin-8 production were assessed from human macrophages with LAM preparations from different species. Our results suggest that the inositol phosphate capping may represent the major cytokine-inducing component of LAMs. This work not only underlines the diversity of LAM structures among various mycobacterial species but also provides new structures that could be useful to dissect the structure-function relationships of these complex molecules.
lipopolysaccharides, antigens, cell wall, structural studies, Mycobacterium, glycolipid, immune response, lipoarabinomannan, tumor necrosis factor, CD1, interleukin-8, lectins, Mycobacterium chelonae
Structure type: homopolymer
Location inside paper: p.30646
Trivial name: LAM, mannan, galactomannan, cell wall heteromannan
Compound class: EPS, O-polysaccharide, cell wall polysaccharide, N-glycan, cell surface polysaccharide, mannan, D-mannan, galactomannan
Contained glycoepitopes: IEDB_130701,IEDB_140116,IEDB_141793,IEDB_141828,IEDB_144983,IEDB_152206,IEDB_153220,IEDB_153762,IEDB_153763,IEDB_76933,IEDB_983930,SB_198,SB_44,SB_67,SB_72
Methods: NMR-2D, NMR, MALDI-MS
Biological activity: tumor necrosis factor-a and interleukin-8 secretion
Comments, role: structural motif of lipoarabinomannan, arabinofuranose chain is attached to the mannose chain of lipoarabinomannan
Related record ID(s): 457, 610, 611, 612
NCBI Taxonomy refs (TaxIDs): 1774,
1773,
33892,
1772Reference(s) to other database(s): GTC:G55317BB, GlycomeDB:
6776, CCSD:
46066, CBank-STR:6345
Show glycosyltransferases
NMR conditions: in DMSO-d6 at 343 K
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6
aDManp 100.7 70.97 72.0 67.0 ? ?
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6
aDManp 4.71 3.68 3.56 3.52 ? ?
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6
aDManp 100.7/4.71 70.97/3.68 72.0/3.56 67.0/3.52 ?/? ?/?
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 |
|---|
| | aDManp | 4.71 | 3.68 | 3.56 | 3.52 | ? | ? |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 |
|---|
| | aDManp | 100.7 | 70.97 | 72.0 | 67.0 | ? | ? |
|
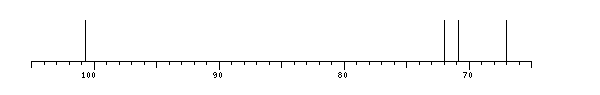
The spectrum also has 2 signals at unknown positions (not plotted). |
There is only one chemically distinct structure:
 report error
report error report error
report error


