Taxonomic group: plant / Streptophyta
(Phylum: Streptophyta)
Organ / tissue: flower
The structure was elucidated in this paperPublication DOI: 10.1016/S0031-9422(97)01044-3Journal NLM ID: 0151434Publisher: Elsevier
Institutions: Chemical Laboratory, Meiji-Gakuin University, Yokohama, Japan, Faculty of Pharmacy, Ankara University, Ankara, Turkey, Faculty of Science, Department of Chemistry, Kocaeli University, Turkey, Laboratory of Floriculture, College of Horticulture, Minami-Kyushu University, Miyazaki, Japan, Institute of Medicinal Chemistry, Hoshi University, Tokyo, Japan
One new flavonol glycoside, kaempferol 3-(4 ",6 "-diacetylglucoside)-7-rhamnoside and three other known kaempferol glycosides: kaempferol 3-glucoside-7-rhamnoside, kaempferol 3-(6 "-acetylglucoside)-7-rhamnoside, and kaempferol 7-rhamnoside, were isolated and characterised from flowers of Delphinium formosum. The structures were elucidated by spectral and chemical methods.
Ranunculaceae, Delphinium formosum, acetylated flavonol glycosides, kaempferol 3-(4, 6 -diacetylglucoside)-7-rhamnoside, kaempferol 3-(6 -acetylglucoside)-7-rhamnoside
Structure type: monomer ; 973 [M+H]+
Location inside paper: p. 244, column 1, paragraph 7, compound 4, table 1, table 2
Compound class: glycoside
Contained glycoepitopes: IEDB_136105,IEDB_225177,IEDB_885823
Methods: 1H NMR, FAB-MS, TLC, acid hydrolysis, HPLC, UV, extraction, acetylation, CC, melting point determination, HMBC, HMQC, COSY, DIFNOE
Related record ID(s): 61901, 61902, 61903, 61905
NCBI Taxonomy refs (TaxIDs): 46246
Show glycosyltransferases
NMR conditions: in DMSO-d6
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9 C10 C11 C12 C13 C14 C15 C16
7 aLRhap 100.1 71.5 72.0 73.4 71.4 18.7
xXKaempferol - 160.8 137.4 177.4 161.9 100.0 163.2 95.7 157.5 106.1 123.3 131.0 116.7 160.8 116.7 131.0
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6 H7 H8 H9 H10 H11 H12 H13 H14 H15 H16
7 aLRhap 5.65 3.5-4.1 3.5-4.1 3.5-4.1 3.5-4.1 1.20
xXKaempferol - - - - - 6.50 - 6.85 - - - 8.17 7.02 - 7.02 8.17
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6 C7/H7 C8/H8 C9/H9 C10/H10 C11/H11 C12/H12 C13/H13 C14/H14 C15/H15 C16/H16
7 aLRhap 100.1/5.65 71.5/3.5-4.1 72.0/3.5-4.1 73.4/3.5-4.1 71.4/3.5-4.1 18.7/1.20
xXKaempferol 100.0/6.50 95.7/6.85 131.0/8.17 116.7/7.02 116.7/7.02 131.0/8.17
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 | H7 | H8 | H9 | H10 | H11 | H12 | H13 | H14 | H15 | H16 |
|---|
| 7 | aLRhap | 5.65 | 3.5
4.1 | 3.5
4.1 | 3.5
4.1 | 3.5
4.1 | 1.20 | |
| | xXKaempferol |
|
|
|
|
| 6.50 |
| 6.85 |
|
|
| 8.17 | 7.02 |
| 7.02 | 8.17 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 | C10 | C11 | C12 | C13 | C14 | C15 | C16 |
|---|
| 7 | aLRhap | 100.1 | 71.5 | 72.0 | 73.4 | 71.4 | 18.7 | |
| | xXKaempferol |
| 160.8 | 137.4 | 177.4 | 161.9 | 100.0 | 163.2 | 95.7 | 157.5 | 106.1 | 123.3 | 131.0 | 116.7 | 160.8 | 116.7 | 131.0 |
|
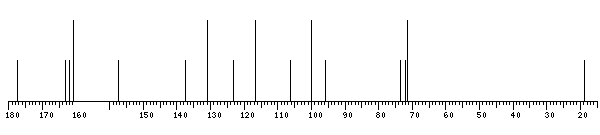
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error report error
report error