Taxonomic group: plant / Streptophyta
(Phylum: Streptophyta)
Organ / tissue: leaf
The structure was elucidated in this paperNCBI PubMed ID: 10514319Publication DOI: 10.1021/np990080oJournal NLM ID: 7906882Publisher: American Society of Pharmacognosy
Correspondence: Pauli GF <pauli

uni-muenster.de>
Institutions: Department of Pharmaceutical Biology, University of Würzburg, Würzburg, Germany, Institute of Pharmaceutical Biology and Phytochemistry, Westfälische Wilhelms-University, Hittorfstrasse 56, D-48149 Münster, Germany
The three major flavonoids isolated from Arabidopsis thaliana plants grown in the greenhouse were identified by means of spectroscopic analysis (UV, NMR, MS) and chiral capillary zone electrophoresis as the novel kaempferol 3-O-β-[β-D-glucopyranosyl(1→6)D-glucopyranoside]-7-O-α-L-rhamnopyranoside (1), kaempferol 3-O-β-D-glucopyranoside-7-O-α-L-rhamnopyranoside (2), and kaempferol 3-O-α-L-rhamnopyranoside-7-O-α-L-rhamnopyranoside (3). Comprehensive NMR studies including selective 1D and gradient-enhanced 2D techniques were applied in order to achieve full signal assignment and definitive proof of linkage for compound 1.
flavonoid, Arabidopsis thaliana, flavone glycoside
Structure type: oligomer ; 755 [M-H]-
C
33H
40O
20Location inside paper: compound 1, table 1, fig. 1
Compound class: flavone glycoside
Contained glycoepitopes: IEDB_136105,IEDB_141806,IEDB_142488,IEDB_146664,IEDB_225177,IEDB_241101,IEDB_885823,IEDB_983931,SB_192
Methods: 13C NMR, 1H NMR, IR, TLC, MALDI-TOF MS, HPLC, UV, capillary electrophoresis (CE), TOCSY, HMBC, COSY, HSQC, HRMS, TFA hydrolysis, reduction with NaCNBH3, API-MS
Comments, role: NMR temperature was not specified
Related record ID(s): 63334, 63335
NCBI Taxonomy refs (TaxIDs): 3702
Show glycosyltransferases
NMR conditions: in vol 95%CD3OD / vol 5%DMSO-d6
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9 C10 C11 C12 C13 C14 C15 C16
3,6 bDGlcp 104.6 75.1 77.8 71.3 77.7 62.5
3 bDGlcp 103.5 75.8 77.9 71.4 77.9 69.5
7 aLRhap 100.0 71.6 72.0 73.5 71.3 18.2
xXKaempferol - 159.3 135.6 179.6 163.3 100.7 163.6 95.9 158.0 107.5 122.5 132.5 116.3 161.8 116.3 132.5
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6 H7 H8 H9 H10 H11 H12 H13 H14 H15 H16
3,6 bDGlcp 4.14 3.04 3.13 3.20 2.97 3.56-3.97
3 bDGlcp 5.35 3.45 3.42 3.35 3.42 3.64-3.98
7 aLRhap 6.58 4.01 3.82 3.47 3.60 1.26
xXKaempferol - - - - - 6.49 - 6.78 - - - 8.14 6.91 - 6.91 8.14
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6 C7/H7 C8/H8 C9/H9 C10/H10 C11/H11 C12/H12 C13/H13 C14/H14 C15/H15 C16/H16
3,6 bDGlcp 104.6/4.14 75.1/3.04 77.8/3.13 71.3/3.20 77.7/2.97 62.5/3.56-3.97
3 bDGlcp 103.5/5.35 75.8/3.45 77.9/3.42 71.4/3.35 77.9/3.42 69.5/3.64-3.98
7 aLRhap 100.0/6.58 71.6/4.01 72.0/3.82 73.5/3.47 71.3/3.60 18.2/1.26
xXKaempferol 100.7/6.49 95.9/6.78 132.5/8.14 116.3/6.91 116.3/6.91 132.5/8.14
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 | H7 | H8 | H9 | H10 | H11 | H12 | H13 | H14 | H15 | H16 |
|---|
| 3,6 | bDGlcp | 4.14 | 3.04 | 3.13 | 3.20 | 2.97 | 3.56
3.97 | |
| 3 | bDGlcp | 5.35 | 3.45 | 3.42 | 3.35 | 3.42 | 3.64
3.98 | |
| 7 | aLRhap | 6.58 | 4.01 | 3.82 | 3.47 | 3.60 | 1.26 | |
| | xXKaempferol |
|
|
|
|
| 6.49 |
| 6.78 |
|
|
| 8.14 | 6.91 |
| 6.91 | 8.14 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 | C10 | C11 | C12 | C13 | C14 | C15 | C16 |
|---|
| 3,6 | bDGlcp | 104.6 | 75.1 | 77.8 | 71.3 | 77.7 | 62.5 | |
| 3 | bDGlcp | 103.5 | 75.8 | 77.9 | 71.4 | 77.9 | 69.5 | |
| 7 | aLRhap | 100.0 | 71.6 | 72.0 | 73.5 | 71.3 | 18.2 | |
| | xXKaempferol |
| 159.3 | 135.6 | 179.6 | 163.3 | 100.7 | 163.6 | 95.9 | 158.0 | 107.5 | 122.5 | 132.5 | 116.3 | 161.8 | 116.3 | 132.5 |
|
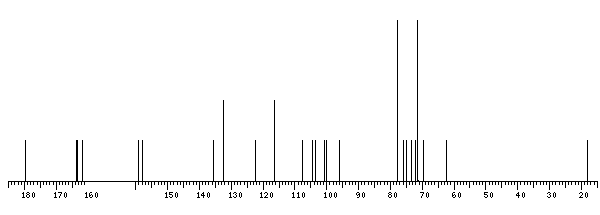
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error Found 1 record.
Displayed record 1
Found 1 record.
Displayed record 1
 report error
report error (previously named: Arabis thaliana)
(previously named: Arabis thaliana) uni-muenster.de>
uni-muenster.de>