Taxonomic group: bacteria / Proteobacteria
(Phylum: Proteobacteria)
The structure was elucidated in this paperNCBI PubMed ID: 15388359Journal NLM ID: 0043535Publisher: Elsevier
Correspondence: evguenii.vinogradov

nrc-cnrc.gc.ca
Institutions: Institute for Biological Sciences, National Research Council, 100 Sussex Dr., Ottawa, ON, Canada K1A 0R6, bUniwersytet Slaski, Katedra Mikrobiologii, ul. Jagiellonska 28, 40-032 Katowice, Poland
The following structure of the Salmonella cerro LPS O-chain repeating unit has been determined using NMR and chemical methods: →4)-α-D-Man(1→2)-α-D-Man(1→2)-β-D-Man(1→3)-α-D-GalN Ac-(1→.
LPS, structure, Salmonella, O-chain, S. cerro
Structure type: polymer chemical repeating unit
Location inside paper: abstract, p.2442
Compound class: O-polysaccharide, O-antigen
Contained glycoepitopes: IEDB_130648,IEDB_130701,IEDB_136104,IEDB_137473,IEDB_137485,IEDB_1391961,IEDB_140116,IEDB_141584,IEDB_141830,IEDB_143632,IEDB_144983,IEDB_152206,IEDB_885822,IEDB_983930,SB_136,SB_196,SB_44,SB_67,SB_72
Methods: methylation, NMR-2D, NMR, sugar analysis
Related record ID(s): 9387, 22712, 25631, 108631, 115468
NCBI Taxonomy refs (TaxIDs): 340188,
590Reference(s) to other database(s): GTC:G82509QD, GlycomeDB:
3519
Show glycosyltransferases
NMR conditions: in D2O at 313 K
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6
3,2,2 aDManp 102.9 71.3 72.1 74.9 72.8 61.4
3,2 aDManp 100.6 79.2 71.2 67.5 73.3 61.4
3 bDManp 102.4 77.0 74.8 68.1 77.9 62.0
2 Ac ? 23.2
aDGalpN 99.4 50.1 77.6 69.6 72.6 62.1
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6
3,2,2 aDManp 5.04 3.99 4.03 3.80 3.78 3.80-3.80
3,2 aDManp 5.33 4.10 4.03 3.79 4.02 3.80-3.80
3 bDManp 4.79 3.99 3.72 3.60 3.38 3.73-3.92
2 Ac - 2.05
aDGalpN 5.31 4.29 3.98 4.17 4.00 3.74-3.74
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6
3,2,2 aDManp 102.9/5.04 71.3/3.99 72.1/4.03 74.9/3.80 72.8/3.78 61.4/3.80-3.80
3,2 aDManp 100.6/5.33 79.2/4.10 71.2/4.03 67.5/3.79 73.3/4.02 61.4/3.80-3.80
3 bDManp 102.4/4.79 77.0/3.99 74.8/3.72 68.1/3.60 77.9/3.38 62.0/3.73-3.92
2 Ac 23.2/2.05
aDGalpN 99.4/5.31 50.1/4.29 77.6/3.98 69.6/4.17 72.6/4.00 62.1/3.74-3.74
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 |
|---|
| 3,2,2 | aDManp | 5.04 | 3.99 | 4.03 | 3.80 | 3.78 | 3.80
3.80 |
| 3,2 | aDManp | 5.33 | 4.10 | 4.03 | 3.79 | 4.02 | 3.80
3.80 |
| 3 | bDManp | 4.79 | 3.99 | 3.72 | 3.60 | 3.38 | 3.73
3.92 |
| 2 | Ac |
| 2.05 | |
| | aDGalpN | 5.31 | 4.29 | 3.98 | 4.17 | 4.00 | 3.74
3.74 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 |
|---|
| 3,2,2 | aDManp | 102.9 | 71.3 | 72.1 | 74.9 | 72.8 | 61.4 |
| 3,2 | aDManp | 100.6 | 79.2 | 71.2 | 67.5 | 73.3 | 61.4 |
| 3 | bDManp | 102.4 | 77.0 | 74.8 | 68.1 | 77.9 | 62.0 |
| 2 | Ac | ? | 23.2 | |
| | aDGalpN | 99.4 | 50.1 | 77.6 | 69.6 | 72.6 | 62.1 |
|
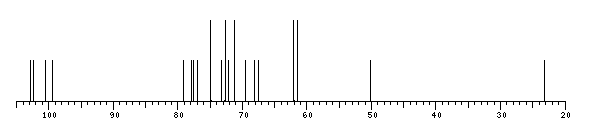
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error report error
report error (previously named: Salmonella cerro K)
(previously named: Salmonella cerro K)

