Taxonomic group: plant, fungi / Streptophyta, Ascomycota
(Phylum: Streptophyta, Ascomycota)
NCBI PubMed ID: 26512632Publication DOI: 10.3390/molecules201019291Journal NLM ID: 100964009Publisher: Basel, Switzerland: MDPI
Correspondence: Qiu ZD <qiuzd

ccucm.edu.cn>; Wang WN <cnweinanwang

163.com>; Yan BX <ybx10528

163.com>; Xu WD <15584452505

163.com>; Qiu Y <ccqy19890810

163.com>; Guo YL <ccyunlong1016

126.com>
Institutions: School of Pharmaceutical Sciences, Changchun University of Chinese Medicine, Changchun, China
Compound K (CK), a highly active and bioavailable derivative obtained from protopanaxadiol ginsenosides, displays a wide variety of pharmacological properties, especially antitumor activity. However, the inadequacy of natural sources limits its application in the pharmaceutical industry. In this study, we firstly discovered that Cordyceps sinensis was a potent biocatalyst for the biotransformation of ginsenoside Rb1 into CK. After a series of investigations on the biotransformation parameters, an optimal composition of the biotransformation culture was found to be lactose, soybean powder and MgSO4 without controlling the pH. Also, an optimum temperature of 30 °C for the biotransformation process was suggested in a range of 25 °C-50 °C. Then, a biotransformation pathway of Rb1→Rd→F2→CK was established using high performance liquid chromatography/quadrupole time-of-flight mass spectrometry (HPLC-Q-TOF-MS). Our results demonstrated that the molar bioconversion rate of Rb1 to CK was more than 82% and the purity of CK produced by C. sinensis under the optimized conditions was more than 91%. In conclusion, the combination of C. sinensis and the optimized conditions is applicable for the industrial preparation of CK for medicinal purposes
optimization, Cordyceps sinensis, biotransformation, compound K, ginsenoside Rb1
Structure type: monomer ; 667.4787 [M+HCOOH−H]−
C
36H
62O
8Location inside paper: Table 1, Rh2, Table 4, Rh2
Trivial name: ginsenoside Rh2
Compound class: glycoside
Contained glycoepitopes: IEDB_142488,IEDB_146664,IEDB_983931,SB_192
Methods: 13C NMR, TLC, extraction, ESI-QTOF-MS, cell growth, HPLC-MS, evaporation, sonication
Synthetic data: enzymatic in vivo
Related record ID(s): 44249, 44478, 47977, 47978, 47979, 47980, 47982, 47983, 47984, 47985, 47986, 47987
NCBI Taxonomy refs (TaxIDs): 4054,
72228
Show glycosyltransferases
NMR conditions: in CD3OD
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9 C10 C11 C12 C13 C14 C15 C16 C17 C18 C19 C20 C21 C22 C23 C24 C25 C26 C27 C28 C29 C30
3 bDGlcp 106.92 ? ? ? ? ?
xXProtopanaxadiol20s 39.12 27.05 88.78 39.66 56.35 18.43 35.15 40.00 50.38 36.94 32.02 70.96 49.54 51.69 31.32 26.70 54.77 16.77 15.61 72.94 26.83 35.88 22.97 126.30 130.73 25.78 17.66 28.14 16.34 17.65
1H NMR data:
missing...
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 | C10 | C11 | C12 | C13 | C14 | C15 | C16 | C17 | C18 | C19 | C20 | C21 | C22 | C23 | C24 | C25 | C26 | C27 | C28 | C29 | C30 |
|---|
| 3 | bDGlcp | 106.92 | ? | ? | ? | ? | ? | |
| | xXProtopanaxadiol20s | 39.12 | 27.05 | 88.78 | 39.66 | 56.35 | 18.43 | 35.15 | 40.00 | 50.38 | 36.94 | 32.02 | 70.96 | 49.54 | 51.69 | 31.32 | 26.70 | 54.77 | 16.77 | 15.61 | 72.94 | 26.83 | 35.88 | 22.97 | 126.30 | 130.73 | 25.78 | 17.66 | 28.14 | 16.34 | 17.65 |
|
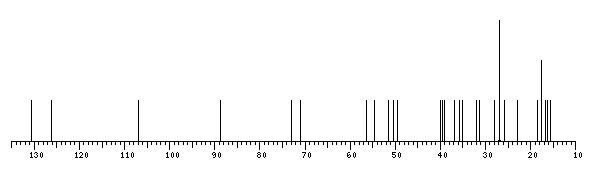
The spectrum also has 5 signals at unknown positions (not plotted). |
There is only one chemically distinct structure:
 report error
report error report error
report error
 (later renamed to: Ophiocordyceps sinensis CICC14017)
(later renamed to: Ophiocordyceps sinensis CICC14017)