Taxonomic group: plant / Streptophyta
(Phylum: Streptophyta)
Publication DOI: 10.1248/cpb.43.338Journal NLM ID: 0377775Publisher: Pharmaceutical Society Of Japan
Institutions: Department of Botany, St. Xavier's College (Autonomous), Palayamkottai, India, Department of Chemistry, Manonmaniam Sundaranar University, Tirunelveli, India, Faculty of Pharmaceutical Sciences, Science University of Tokyo, Tokio, Japan
Three dihydroflavonol glycosides, 3-O-α-L-rhamnosides of (2S,3S)-, (2R,3R)- and (2R,3S)-5,7,3',4'-tetrahydroxydihydroflavonol, were isolated from Sphaerostephanos arbuscula as the first example of dihydroflavonols from ferns. Aglycone chirality was determined by 1H-NMR data.
1H-NMR, astilbin, Sphaerostephanos arbuscula, dihydroflavonol, chirality determination, fern
Structure type: oligomer
C
27H
30O
16Location inside paper: p. 339, rutin
Trivial name: rutin, rutoside, rutin, quercetin rutinoside, rutoside, quercetin 3-O-rutinose, quercetin-3-O-rutinoside, quercetin 3-O-rutinoside
Compound class: saponin glycoside, glycoside, flavonoid glycoside, flavonol glycoside, flavone glycoside
Contained glycoepitopes: IEDB_136105,IEDB_142488,IEDB_144144,IEDB_146664,IEDB_225177,IEDB_885823,IEDB_983931,SB_192
Methods: 13C NMR, 1H NMR, EI-MS, IR, acid hydrolysis, GLC, HPLC, UV, extraction, optical rotation measurement, CD, CC, melting point determination, precipitation, derivatization
Related record ID(s): 50488, 50489, 50490
NCBI Taxonomy refs (TaxIDs): 1521241Reference(s) to other database(s): CCSD:
50720, CBank-STR:3399
Show glycosyltransferases
NMR conditions: in DMSO-d6
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9 C10 C11 C12 C13 C14 C15 C16
3,6 aLRhap 100.7 70.4 70.0 71.8 68.2 17.7
3 bDGlcp 101.2 74.0 76.4 70.6 75.9 67.0
xXQuercetin - 156.4 133.3 177.4 161.2 98.7 164.1 93.6 156.6 104.0 121.2 115.2 144.7 148.4 116.3 121.6
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6 H7 H8 H9 H10 H11 H12 H13 H14 H15 H16
3,6 aLRhap 4.40 ? ? ? ? 1.00
3 bDGlcp 5.35 ? ? ? ? ?
xXQuercetin - - - - - 6.20 - 6.39 - - - 6.85 - - ? ?
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6 C7/H7 C8/H8 C9/H9 C10/H10 C11/H11 C12/H12 C13/H13 C14/H14 C15/H15 C16/H16
3,6 aLRhap 100.7/4.40 70.4/? 70.0/? 71.8/? 68.2/? 17.7/1.00
3 bDGlcp 101.2/5.35 74.0/? 76.4/? 70.6/? 75.9/? 67.0/?
xXQuercetin 98.7/6.20 93.6/6.39 115.2/6.85 116.3/? 121.6/?
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 | H7 | H8 | H9 | H10 | H11 | H12 | H13 | H14 | H15 | H16 |
|---|
| 3,6 | aLRhap | 4.40 | ? | ? | ? | ? | 1.00 | |
| 3 | bDGlcp | 5.35 | ? | ? | ? | ? | ? | |
| | xXQuercetin |
|
|
|
|
| 6.20 |
| 6.39 |
|
|
| 6.85 |
|
| ? | ? |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 | C10 | C11 | C12 | C13 | C14 | C15 | C16 |
|---|
| 3,6 | aLRhap | 100.7 | 70.4 | 70.0 | 71.8 | 68.2 | 17.7 | |
| 3 | bDGlcp | 101.2 | 74.0 | 76.4 | 70.6 | 75.9 | 67.0 | |
| | xXQuercetin |
| 156.4 | 133.3 | 177.4 | 161.2 | 98.7 | 164.1 | 93.6 | 156.6 | 104.0 | 121.2 | 115.2 | 144.7 | 148.4 | 116.3 | 121.6 |
|
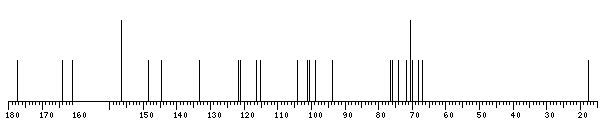
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error report error
report error