Taxonomic group: plant / Streptophyta
(Phylum: Streptophyta)
Organ / tissue: flower
The structure was elucidated in this paperNCBI PubMed ID: 10444863Publication DOI: 10.1016/S0031-9422(99)00109-0Journal NLM ID: 0151434Publisher: Elsevier
Institutions: Chemistry Department, Royal Veterinary and Agricultural University, Thorvaldsensvej 40, DK-1871 Frederiksberg C, Denmark, Chemical Instrument Center, Nagoya University, Chikusa, Nagoya 464-8602, Japan
From the flower extracts of Crocus speciosus and C. antalyensis nine flavonol glycosides have been isolated. One of these products is a new flavonol glycoside identified as kaempferol 3-O-α-(2,3-di-O-β-D-glucopyranosyl)rhamnopyranoside by UV, mass and NMR spectroscopy.
1H, 13C, 2D NMR, flavonol glycosides, Iridaceae, Crocus speciosus, C. antalyensis, kaempferol 3-O-α-(2, 3-di-O-β-D-glucopyranosyl)rhamnopyranoside
Structure type: monomer ; 449 [M+H]+
C
21H
20O
11Location inside paper: compound 9, fig. 1(9), table 1(9), table 2(9), table 3(9)
Compound class: flavonol glycoside
Contained glycoepitopes: IEDB_142488,IEDB_146664,IEDB_983931,SB_192
Methods: 13C NMR, 1H NMR, FAB-MS, HPLC, UV, HMBC, COSY, HOHAHA, HSQC
Comments, role: NMR temperature was not specified
Related record ID(s): 63955, 63956, 63957, 63958, 63959, 63960, 63961, 63962
NCBI Taxonomy refs (TaxIDs): 481018
Show glycosyltransferases
NMR conditions: in vol 90%DMSO-d6 / vol 10%CF3COOD
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9 C10 C11 C12 C13 C14 C15 C16
3 bDGlcp 101.2 74.0 76.1 69.6 77.0 60.6
xXKaempferol - 156.4 133.3 177.4 161.0 98.4 163.9 93.4 156.4 104.0 120.9 130.7 117.5 159.8 117.5 130.7
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6 H7 H8 H9 H10 H11 H12 H13 H14 H15 H16
3 bDGlcp 5.39 3.10 3.21 3.13 3.24 3.34-3.54
xXKaempferol - - - - - 6.18 - 6.38 - - - 7.98 6.85 - 6.85 7.98
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6 C7/H7 C8/H8 C9/H9 C10/H10 C11/H11 C12/H12 C13/H13 C14/H14 C15/H15 C16/H16
3 bDGlcp 101.2/5.39 74.0/3.10 76.1/3.21 69.6/3.13 77.0/3.24 60.6/3.34-3.54
xXKaempferol 98.4/6.18 93.4/6.38 130.7/7.98 117.5/6.85 117.5/6.85 130.7/7.98
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 | H7 | H8 | H9 | H10 | H11 | H12 | H13 | H14 | H15 | H16 |
|---|
| 3 | bDGlcp | 5.39 | 3.10 | 3.21 | 3.13 | 3.24 | 3.34
3.54 | |
| | xXKaempferol |
|
|
|
|
| 6.18 |
| 6.38 |
|
|
| 7.98 | 6.85 |
| 6.85 | 7.98 |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 | C10 | C11 | C12 | C13 | C14 | C15 | C16 |
|---|
| 3 | bDGlcp | 101.2 | 74.0 | 76.1 | 69.6 | 77.0 | 60.6 | |
| | xXKaempferol |
| 156.4 | 133.3 | 177.4 | 161.0 | 98.4 | 163.9 | 93.4 | 156.4 | 104.0 | 120.9 | 130.7 | 117.5 | 159.8 | 117.5 | 130.7 |
|
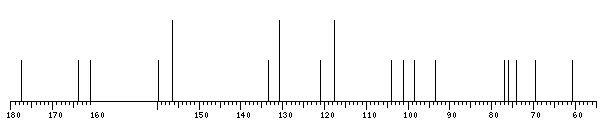
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error report error
report error