Taxonomic group: bacteria / Proteobacteria
(Phylum: Proteobacteria)
Associated disease: infection due to Escherichia coli [ICD11:
XN6P4 
]
The structure was elucidated in this paperNCBI PubMed ID: 7544238Publication DOI: 10.1016/0008-6215(95)00041-QJournal NLM ID: 0043535Publisher: Elsevier
Correspondence: torgov

ioc.ac.ru
Institutions: Max-Planck-Institut für Immunbiologie, Freiburg, Germany
Structures for the N-acetylneuraminic acid (Neu5Ac)-containing O56 and O24 polysaccharides of Escherichia coli have been reported previously. During these studies unusual chemical shifts had been observed for the NMR signals for H-3eq and C-3 of the Neu5Ac residues of both polysaccharides. In further pursuing this phenomenon, we have reinvestigated the O56 and O24 polysaccharides as well as derived oligosaccharides by one- and two-dimensional NMR spectroscopy. The results showed that structures of both polysaccharides (PSs) had to be modified and formulated as [formula: see text] 2D ROESY spectra revealed a strong NOE between H-3eq of Neu5Ac and the protons of the side-chain sugar (H-3 and H-5 of α-D-Galp in the O56 PS and H-3 of α-D-Glcp in the O24 PS) and also between H-3ax of Neu5Ac and H-3 of β-D-Glcp in the main chain. This indicated a close spatial association of the seven-linked α-Neu5Ac and the side-chain residues α-D-Galp (O56 PS) and α-D-Glcp (O25 PS), respectively. The strong long-range spatial contacts caused the unusual chemical shifts of H-3eq and C-3 of Neu5Ac.
Escherichia coli, NMR spectroscopy, polysaccharide structure, 024 and 056 antigens
Structure type: suggested polymer biological repeating unit
Location inside paper: abstract, structure V, Table 8
Compound class: O-polysaccharide, O-antigen
Contained glycoepitopes: IEDB_135813,IEDB_136794,IEDB_136906,IEDB_137340,IEDB_137472,IEDB_141794,IEDB_141807,IEDB_142488,IEDB_146100,IEDB_146664,IEDB_149174,IEDB_151528,IEDB_151531,IEDB_190606,IEDB_983931,SB_170,SB_171,SB_172,SB_192,SB_7,SB_84
Methods: NMR-2D, NMR, sugar analysis, Smith degradation
Comments, role: revision of structure ID 117052; chemical repeat frame is different in the paper; carbon 5 in aXNeup at #3,3 has a +15.4 ppm mismatch in 13C spectra (experiment: 69.6; hybrid simulation in water: 54.2, trust 60%)
Related record ID(s): 326, 327, 328, 8382, 8411, 8412, 8413, 8414, 20438, 20659, 22619, 25691, 108697, 124618
NCBI Taxonomy refs (TaxIDs): 562Reference(s) to other database(s): GTC:G32692RT, GlycomeDB:
6888
Show glycosyltransferases
NMR conditions: in D2O
[as TSV]
13C NMR data:
Linkage Residue C1 C2 C3 C4 C5 C6 C7 C8 C9
3,3,5 Ac
3,3 aXNeup 173.9 102.4 37.75 69.6 ? 72.5 78.9 73.0 63.7
3,2 aDGalp 97.2 69.6 70.6 70.5 71.55 62.4
3 bDGlcp 103.3 73.6 78.7 69.9 76.75 62.4
2 Ac
bDGlcpN 102.2 57.3 80.8 70.6 76.3 62.45
1H NMR data:
Linkage Residue H1 H2 H3 H4 H5 H6 H7 H8 H9
3,3,5 Ac
3,3 aXNeup - - 2.0-2.51 4.05 3.91 3.88 3.72 3.78 3.50-3.80
3,2 aDGalp 5.52 3.83 3.92 4.00 4.13 3.75
3 bDGlcp 4.70 3.66 4.15 3.54 3.50 3.73-3.90
2 Ac
bDGlcpN 4.74 3.68 4.08 3.52 3.53 3.79-4.00
1H/13C HSQC data:
Linkage Residue C1/H1 C2/H2 C3/H3 C4/H4 C5/H5 C6/H6 C7/H7 C8/H8 C9/H9
3,3,5 Ac
3,3 aXNeup 37.75/2.0-2.51 69.6/4.05 ?/3.91 72.5/3.88 78.9/3.72 73.0/3.78 63.7/3.50-3.80
3,2 aDGalp 97.2/5.52 69.6/3.83 70.6/3.92 70.5/4.00 71.55/4.13 62.4/3.75
3 bDGlcp 103.3/4.70 73.6/3.66 78.7/4.15 69.9/3.54 76.75/3.50 62.4/3.73-3.90
2 Ac
bDGlcpN 102.2/4.74 57.3/3.68 80.8/4.08 70.6/3.52 76.3/3.53 62.45/3.79-4.00
1H NMR data:
| Linkage | Residue | H1 | H2 | H3 | H4 | H5 | H6 | H7 | H8 | H9 |
|---|
| 3,3,5 | Ac | |
| 3,3 | aXNeup |
|
| 2.0
2.51 | 4.05 | 3.91 | 3.88 | 3.72 | 3.78 | 3.50
3.80 |
| 3,2 | aDGalp | 5.52 | 3.83 | 3.92 | 4.00 | 4.13 | 3.75 | |
| 3 | bDGlcp | 4.70 | 3.66 | 4.15 | 3.54 | 3.50 | 3.73
3.90 | |
| 2 | Ac | |
| | bDGlcpN | 4.74 | 3.68 | 4.08 | 3.52 | 3.53 | 3.79
4.00 | |
|
13C NMR data:
| Linkage | Residue | C1 | C2 | C3 | C4 | C5 | C6 | C7 | C8 | C9 |
|---|
| 3,3,5 | Ac | |
| 3,3 | aXNeup | 173.9 | 102.4 | 37.75 | 69.6 | ? | 72.5 | 78.9 | 73.0 | 63.7 |
| 3,2 | aDGalp | 97.2 | 69.6 | 70.6 | 70.5 | 71.55 | 62.4 | |
| 3 | bDGlcp | 103.3 | 73.6 | 78.7 | 69.9 | 76.75 | 62.4 | |
| 2 | Ac | |
| | bDGlcpN | 102.2 | 57.3 | 80.8 | 70.6 | 76.3 | 62.45 | |
|
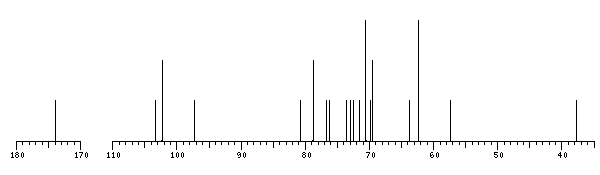
The spectrum also has 1 signal at unknown position (not plotted). |
There is only one chemically distinct structure:
 report error
report error Found 1 record.
Displayed record 1
Found 1 record.
Displayed record 1
 report error
report error
 ]
] ioc.ac.ru
ioc.ac.ru